热门话题
#
Bonk 生态迷因币展现强韧势头
#
有消息称 Pump.fun 计划 40 亿估值发币,引发市场猜测
#
Solana 新代币发射平台 Boop.Fun 风头正劲
计算机学习RNA碱基配对规则需要什么?
人们正在训练大型语言模型以预测RNA结构。这些模型中的一些具有数亿个参数。
一个令人兴奋的早期结果是,这些模型直接从数据中学习了沃森-克里克-富兰克林碱基配对的规则。
哈佛大学的一个研究小组决定看看能够实现这一结果的最小模型是什么。
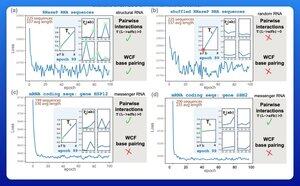
他们使用梯度下降训练了一个只有21个参数的小型概率模型。
仅用50个RNA序列——没有相应的结构——碱基配对的规则在仅仅几个训练周期后就会显现出来。
因此,他们最初问题的答案是,学习这种类型的模型“远比你想象的要少得多”。
我认为这并不意味着大规模训练工作必然是愚蠢或误导的。但这个结果表明,架构创新仍然可以挖掘出很多效率和性能。
生物学语言中有很多潜在的结构。
热门
排行
收藏













